Deep Learning and Bioengineered Feature Engineering for Automated Taxonomic Identification of Mediterranean and Atlantic Demospongiae
DOI:
https://doi.org/10.63318/waujpasv4i1_33Keywords:
Demospongiae taxonomy, Imbalanced classification, Bioengineered features, Deep learning, Marine biodiversity, hierarchical modeling, UCI sponge datasetAbstract
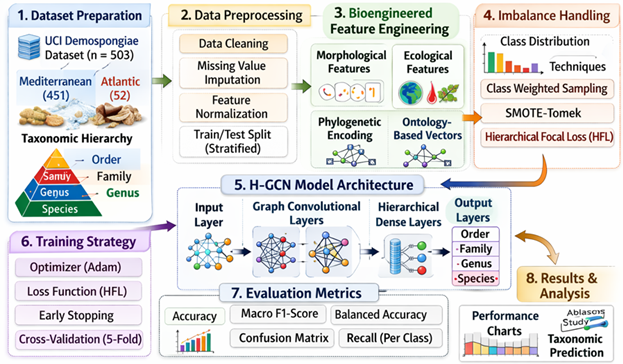
Given morphological convergence, cryptic speciation, and severe class imbalance in regional collection records, accurate taxonomic dentification of marine Demospongiae remains a major challenge in biodiversity monitoring. This study presents a novel computational framework that integrates imbalanced multi-class deep learning with biologically informed feature engineering to automate the hierarchical classification of 503 Demospongiae specimens collected from Mediterranean and Atlantic habitats. We introduce a biologically grounded feature construction pipeline that encodes morphological, ecological, and evolutionary priors derived from sponge ontology structures, generating high-dimensional representations compatible with deep neural architectures. To address strong distributional skew across 7 orders, 42 families, 114 genera, and 230 species, we implement a hybrid imbalance mitigation strategy combining topology-aware synthetic minority oversampling, adaptive class-weighted sampling, and hierarchical focal loss. The proposed architecture further employs a multi-task graph convolutional network to jointly learn taxonomic relationships while preserving hierarchical constraints. Experimental evaluation demonstrates substantial improvements over conventional machine learning baselines, achieving macro-averaged F1-scores of 0.89 at the order level and 0.76 at the species level, with notable gains in recall for underrepresented Atlantic taxa. Ablation analyses further indicate that the incorporation of bioengineered features significantly enhances model generalization.
Downloads
Downloads
Published
Issue
Section
License
Copyright (c) 2026 The author

This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License.
This journal uses Creative Commons Attribution-Noncommerical 4.0 International License (CC BY-NC 4.0), which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. To view a copy of this license, visit https://creativecommons.org/licenses/by-nc/4.0/.
Copyright of articles
Authors retain copyright of their articles published in this journal.




